|
END is a specially pattern that matches after all lines are processed.This code computes the occurrences of all columns, and prints a sorted report for each of them: # columnvalues.  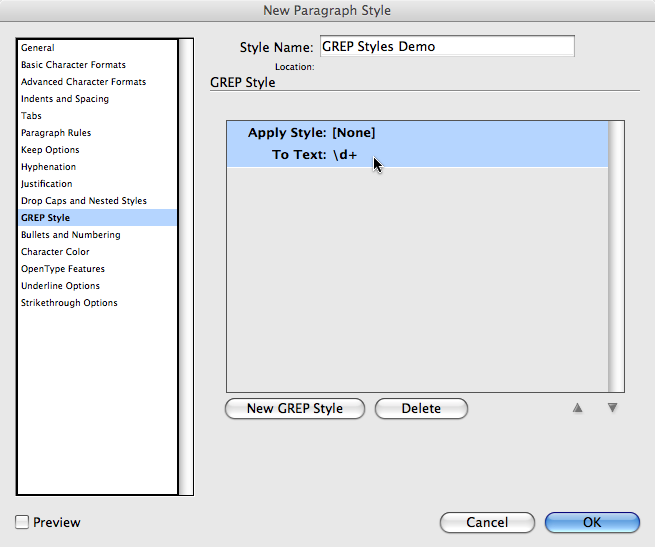 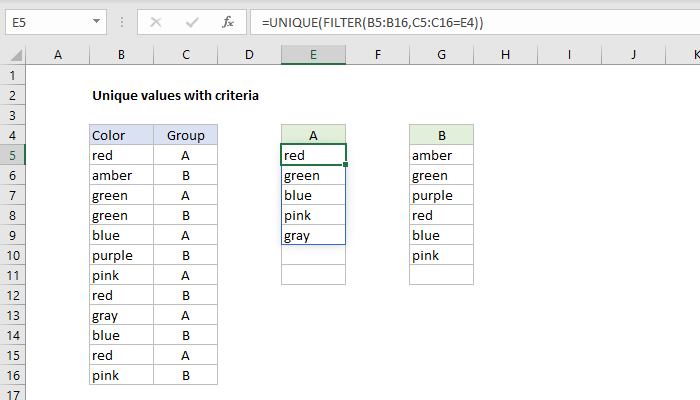
| awk -F, '$2 = "val1" & $3 = "val2" 'Īwk automatically trims whitespace from the values. When i do: grep foo, I get all the lines that contain foo. And the output that you get from all of these commands is: Bar Baz Foo. So, the three following commands all accomplish the same thing: cat test sort sort < test sort test. I have a log file with multiple occurrences of 'foo' for instance. sort can operate either on STDIN redirection, the input from a pipe, or, in the case of a file, you also can just specify the file on the command. If awk is available (it is pretty standard) you can do it like this: $ echo -e 'a,b, c,d\na,val1,val2,c' \ How do i grep unique occurrences from a file. Thats just like passing patterns on the command line (with the -e option if theres more than one), except that when youre calling from a shell you may need to quote the pattern to protect special characters in it from being expanded by the shell. Extract unique values (including first duplicates) by using Advanced Filter. Another option to grep two strings: grep 'word1 \ word2' input. The -f option specifies a file where grep reads patterns. Finally, try on older Unix shells/oses: grep -e pattern1 -e pattern2. Next use extended regular expressions: grep -E 'pattern1pattern2'. I would like to read every uniq value from Column A with only the first hit from column B. The pipe grep "Value1" | grep "Value2" from you question does not do what you specify - it would match too much, e.g.: The syntax is: Use single quotes in the pattern: grep 'pattern' file1 file2. Two columns A and B: Column A have repeated ids and column B has different values for each corresponding repeated value. uniq filters out the adjacent matching lines from the input file (that is. In simple words, uniq is the tool that helps to detect the adjacent duplicate lines and also deletes the duplicate lines. If the contents of 'missing.txt' are fixed strings, not regular expressions, this will speed up the process: grep -F -f missing.txt file.txt > results.txt. Please take note that the column indexing in csvfilter starts with 0 (unlike awk, which starts with 1). If you have python (and you should), you can install it simply like this: pip install csvfilter. grep -f missing.txt file.txt > results.txt. It works like cut, but properly handles CSV column quoting: csvfilter -f 1,3,5 in.csv > out.csv. As you can see in the man page, option -u stands for 'unique', -t for the field delimiter, and -k allows you to select the location (key). Note: As soon as the shell sees the for command it will know it is a. The meta-characters ^ and $ match the beginning and the end of a line. The uniq command in Linux is a command-line utility that reports or filters out the repeated lines in a file. Use greps -f option: Then you only need a single grep call and no loop. You can use simply the options provided by the sort command: sort -u -t, -k4,4 file.csv. grep ACGT creatures/ cut -d : -f 1 sed s/creatures// /g uniq. Note that the grep pattern allows trailing whitespace after/before the values (you can remove the * sub-patterns if you don't want that). For testing purposes, the columns 2 and 3 are used instead of 14 and 15. To get the number of occurrences of each unique value, use uniqs -c option: sort mylist. 
| cut -d ',' -f2,3 | grep '^ *val1 *, *val2 *$' | wc -lĪssuming, as delimiter (and no somehow escaped, is included) in the input. That will give you the number of unique values. If awk is not available you can do it with cut, grep and wc: $ echo -e 'a,b, c,d\na,val1 ,val2,c' \
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
